Interactive Visualization of Molecular Orbitals
Description
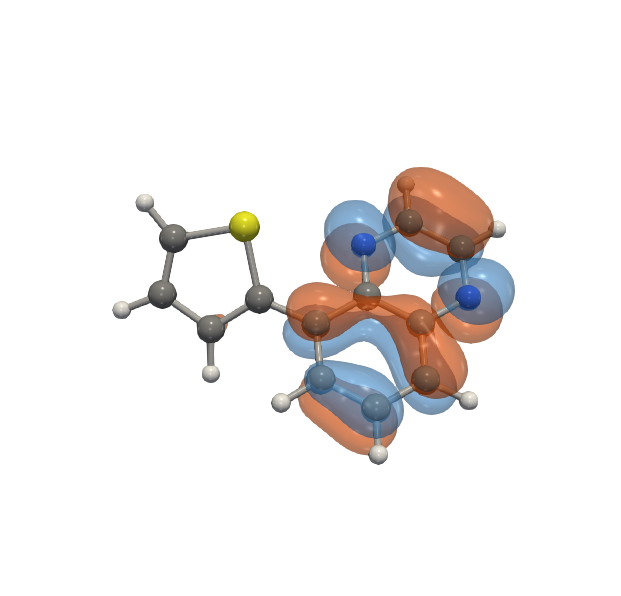
Molecular orbitals specify the spatial distribution of electrons in a molecule. Using software tools like VeloxChem, these orbitals can be computed as a scalar function on a grid enclosing the molecule. Isosurface visualization is the standard method of visualizing these orbitals. However, currently the computation of the volumetric representaion of orbitals is slow which does not allow for interactive visualization.
The goal of this project is to implement a tool for intercative 3D visualization of molecular orbitals. Achieving interactivity will require implementaion of a faster approximate molecular orbital computation algorithm which has been developed in-house. This tool is meant to be used by theoretical chemists who are more familiar with Python-based tools and interfaces. So, our aim will be to develop a visualizer compatible with Jupyter Notebook. This project will require familiarity with Python in addition to knowledge of C++ and OpenGL.
Useful links
- High performance computation and interactive display of molecular orbitals on GPUs and multi-core CPUs. https://dl.acm.org/doi/pdf/10.1145/1513895.1513897
- https://veloxchem.org/docs/intro.html
- https://jupyter.org/